Sequencing Education

Sequencing is the process of reading the nucleotides present in DNA or RNA molecules. There are two types of sequencing technologies that are used today: Sanger sequencing and next-generation sequencing.
Our scientists have long been at the forefront of sequencing technology, starting with the introduction of the first capillary sequencers over 25 years ago, continuing with advances in Sanger sequencing, and then establishing a new paradigm in next-generation sequencing (NGS) with Ion Torrent™ semiconductor sequencing. Our innovative research technologies are currently paving the way for faster and more affordable sequencing applications in targeted DNA and RNA sequencing, exome sequencing, and genome sequencing.
Sequencing is the ultimate genetic analysis technology for the 21st century
Select an option to determine the best platform for your research needs:
I want to learn more about sequencing and choose the right technology for me.
I want to learn about NGS and how it can enhance my Sanger sequencing efforts.
I want to learn about NGS and how to transition from microarrays.
Sequencing overview
As soon as Watson and Crick discovered the structure of DNA and helped introduce the “central dogma” of biology, the next big genetic challenge became clear: sequence the entire human genome and finally see our genetic blueprint. This milestone was accomplished successfully in 2001, through the Human Genome Project, a global collaborative effort that spanned 10 years and was supported by thousands of scientists and billions of dollars.
In 2008, American scientist Dr. Jonathan Rothberg sequenced a genome—James Watson's—in four months for just $1.5 million dollars. He used a method that came to be called next-generation sequencing, or NGS. It was a completely different approach from traditional sequencing methods that made the promise of the $1000 genome more attainable than ever before.
Today, both Sanger sequencing and NGS are critical methods shaping our understanding of many scientific disciplines, including clinical research and genetic, environmental, and forensic sciences.
The history of sequencing
The development of Sanger sequencing
DNA sequencing was first carried out in the 1970s. British biochemist Dr. Frederick Sanger developed a method of sequencing DNA that has become the gold standard. This method involves copying single-stranded DNA using chemically altered bases, called dideoxynucleotides, that cause the DNA chain to terminate when these bases are incorporated. The terminated chains are then resolved by capillary electrophoresis.
Initially, Sanger sequencing’s throughput was only tens to hundreds of bases per day, but by the mid-1980s the first commercial DNA sequencers (from Applied Biosystems) based on the Sanger sequencing method were producing thousands of bases of sequence a day. By automating the data collection (up to that time the base sequences had to be input into a computer by hand), the accuracy and throughput of DNA sequencing grew enormously. By the mid-1990s, DNA sequencers could be found in many laboratories and were producing as many as a million bases, or a megabase, of sequence per day.
The Human Genome Project (1990-2001)
Sanger-based DNA sequencers are the machines that supported the Human Genome Project; the technology is reliable and continues to be widely used. A form of Sanger sequencing called “shotgun” sequencing was adopted for the Human Genome Project. Also developed by Sanger, the shotgun method is a faster way to analyze an entire genome. While traditional Sanger sequencing analyzes long reads (like those in a whole genome) by “walking” through the chromosome base pair by base pair, shotgun sequencing takes a more random approach. Here, DNA is broken up into small random segments, which are sequenced using the Sanger chain termination method. Several rounds of fragmentation and sequencing are needed to produce overlapping reads for the target DNA. Computer models then help assemble the overlapping reads into a sequence.

The completion of the Human Genome Project led to the discovery of thousands of disease-related genes and the development of genetic tests to help in the diagnosis and determination of genetic predisposition to human disease, and also provided new avenues for advances in biotechnology and drug development.
Next-generation sequencing-searching for scale
After the completion of the Human Genome Project, DNA sequencers were used routinely in laboratories around the world to study genetic variation, and were capable of producing ~2 Mb of sequence per day. While this throughput was sufficient for many studies, sequencing a single human genome would still require years using the Sanger sequencing method.

Starting in 2005, a handful of companies introduced what is now called “next-generation sequencing” (NGS). Unlike the Sanger sequencing method that uses chain termination with dideoxynucleotides, these new technologies created a massively parallel, miniaturized, solid-phase method of sequencing on a single substrate. This method of sequencing was based on a "sequencing by synthesis" principle, reading individual bases as they grow along a polymerized strand, unlike the Sanger method that separates out dideoxynucleotide-terminated reaction products.
Learn more about Ion Torrent™ next-generation sequencing technology ›
Video: Introduction to Ion Torrent™ Next-Generation Sequencing Technology
Ion Torrent™ technology directly translates chemically encoded information (A, C, G, T) into digital information (0, 1) on a semiconductor chip. This approach marries simple chemistry to proprietary semiconductor technology. The result is a sequencing technology that is simpler, faster, and more cost effective and scalable than any other next-generation sequencing technology available.
Next-generation sequencing is capable of generating gigabases of sequence per day, and this massive increase in capacity has led to high-throughput sequencing applications that can be accomplished in a fraction of the time and for a fraction of the cost of previous methods. It has also enabled new kinds of experiments that were not feasible before, such as cDNA sequencing (also called RNA-Seq) that employs tens of millions of sequence reads per sample in emerging research applications such as analysis of alternative splice variants, fusion transcripts, and allele-specific expression.
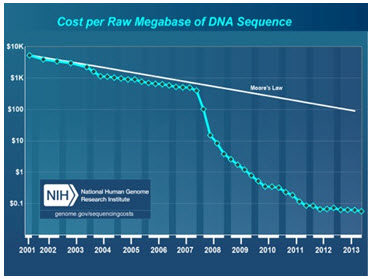
Figure 1: Dramatic reduction in DNA sequencing costs resulting from the massive throughput increase enabled by NGS (data from the NHGRI Genome Sequencing Program).

 CONTACT US
CONTACT US











